atEPDnew, the Arabidopsis thaliana (thale cress) curated promoter database
| Version |
004 |
| Coverage |
22703 promoters 22701
genes |
| Genome assembly |
A. thaliana (Feb 2011 TAIR10/araTha1) |
| Gene annotation |
TAIR 10 genes (1-Feb-2015) |
| Based on data from |
PEAT data from Morton et al. 2014
DeepCAGE from Cumbie et al. 2015
CAGE and OligoCap from Tokizawa et al. 2017
CAGE from Ushijima et al. 2017
EPD (old) |
| Documentation & Viewer(s) |
Promoter assembly pipeline description
EPD viewer
(track content)
Araport (araTha1) |
|
|
Promoter Selection and Anaysis tools
Various tools allow you to analyse promoters from EPD and/or to
select subsets of promoters. In order to analyze the complete EPD
promoter set, go directly to one of the analysis pages. If you
prefer to first select a subset of promoters, go to one of the
selection pages. From the output of the selection pages you can then
directly navigate to one of the analyses pages, or you can continue
with another selection page to refine your promoter selection.
Selection tools
-
EPD
selection tool: Promoter subset selection based on
EPD-supplied annotation.
-
ChIP-Cor: Promoter subset selection based on
experimental data or genome annotations residing in the MGA
repository. Example: select promoters that have more than
100 H3K4me3 ChIP-seq tags data between -100 and +100
relative to the TSS.
-
FindM: Promoter subset selection based on DNA motif
occurrences. Example: select promoters that have (or do not have)
a c-Myc binding site between -100 and +100 relative to
the TSS.
|
Analysis tools
-
ChIP-Cor
: Generation of an aggregation plot (feature
correlation plot) for a specific chromatin of genome
annotation features. Example: Distribution of nucleosomes
(MNase-seq tags) near promoters,
e.g. from -1000 to +1000 relative to the TSS.
-
ChIP-Extract : Extraction of
specific chromatin features around each promoter in
table format. The output is a table with rows
representing each promoter and columns the feature
tag occurance at a specific distance. Example:
Distribution of nucleosomes (MNase-seq tags) near
each promoter, e.g. from -1000 to +1000 relative
to the TSS. Useful for downstream analysis in R,
for example to classify promoters according to
differences in feature distribution.
-
OProf
: Generate a motif occurrence profile around
TSS positions. Example: Generate a plot showing the
occurrence frequency of TATA boxes between -100 to +100
relative to the TSS.
-
FindM
: Extract DNA motif positions near transcription start
sites. Example: extract coordinates of CCAAT boxes
located between -150 and -50 relative to a TSS. The output
is a set of CCAAT-box positions that can be further analyzed
in the same way as a set of TSS positions.
|
|
How-To Documentation:
OProf,
FindM and
ChIP-Cor.
|
Database quality control
Core promoter elements' enrichment
Core promoter element analysis is performed in order to investigate
the quality of the promoter collection. It leverages the preferential
occurrence of certain DNA motifs at characteristic distances from the
TSS. For instance, TATA boxes occur in a narrow region
centered about 28 bp upstream of the TSS, whereas the CCAAT box
occurs in a much wider area, with a maximal frequency at position
-80. Based on these observations, a high-quality promoter collection
is expected to show high peaks for both motifs. In addition, a narrow
TATA box peak at -28 would indicate precise TSS mapping. This analysis
has been performed using
OProf. EPD users are
encouraged to repeat this analysis and to perform others in order to
check the quality of the promoter list.
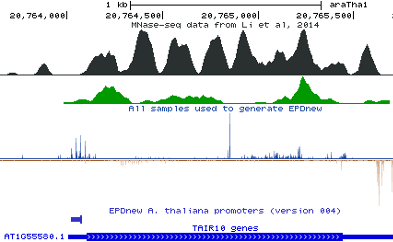
TAIR 10 genes (1-Feb-2015)
PEAT data from Morton et al. 2014;DeepCAGE from Cumbie et al. 2015;CAGE and OligoCap from Tokizawa et al. 2017;CAGE from Ushijima et al. 2017;EPD (old)
chr1%3A20762776-20766776
AT1G55580_1
TATA-box: this core promoter element is normally found 28
bp upstream the transcription start site. The following plot shows
that EPDnew promoter collection has a more focused TATA-box
distribution compared to TAIR10 annotation suggesting a precise TSS
mapping in EPDnew.
Initiator: it is found at the TSS and shows a great
enrichemnt in EPDnew compared to TAIR10 promoter collection.